Zika virus structure revealed
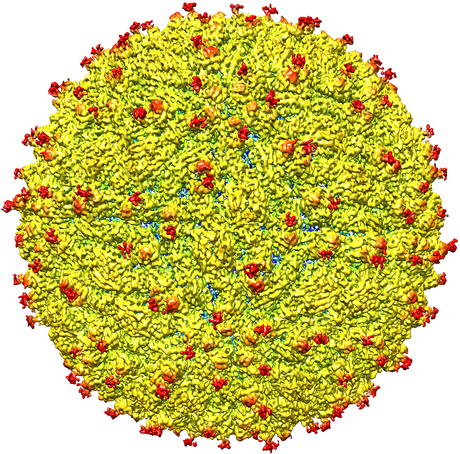
A research team led by Purdue University has determined the structure of the Zika virus, revealing insights critical to the development of effective antiviral treatments and vaccines. The team also identified regions within the virus structure where it differs from other flaviviruses — the family of viruses to which Zika belongs.
In the majority of individuals infected with Zika virus, symptoms are mild and include fever, skin rashes and flu-like illness. However, the mosquito-borne disease has recently been associated with a birth defect called microcephaly that causes brain damage and an abnormally small head in babies born to mothers infected during pregnancy. It also has been associated with the autoimmune disease Guillain-Barré syndrome, which can lead to temporary paralysis.
A Purdue research team led by Richard Kuhn and Michael Rossmann used cryo-electron microscopy to determine the virus structure at a resolution that previously would only have been possible through X-ray crystallography. Rossmann explained that X-ray crystallography is the standard method for determining the structure of viruses, but it requires a relatively large amount of virus, which isn’t always available and it takes a long time.
The team studied a strain of Zika virus isolated from a patient infected during the French Polynesia epidemic and determined the structure to 3.8 Å. At this near-atomic resolution, key features of the virus structure can be seen and groups of atoms that form specific chemical entities, such as those that represent one of 20 naturally occurring amino acids, can be recognised, Rossmann said.
“The structure of the virus provides a map that shows potential regions of the virus that could be targeted by a therapeutic treatment, used to create an effective vaccine or to improve our ability to diagnose and distinguish Zika infection from that of other related viruses,” said Kuhn. “Determining the structure greatly advances our understanding of Zika — a virus about which little is known.”
Writing in the journal Science, the team revealed that the virus structure was very similar to that of other flaviviruses — such as dengue, West Nile, yellow fever, Japanese encephalitis and tickborne encephalitic viruses — with an RNA genome surrounded by a lipid membrane inside an icosahedral protein shell. Any regions within the virus structure unique to Zika have the potential to explain differences in how a virus is transmitted and how it manifests as a disease, said Kuhn.
“Most viruses don’t invade the nervous system or the developing foetus due to blood-brain and placental barriers, but the association with improper brain development in foetuses suggests Zika does,” said Purdue graduate student Devika Sirohi. “It is not clear how Zika gains access to these cells and infects them, but these areas of structural difference may be involved. These unique areas may be crucial and warrant further investigation.”

The team found that all of the known flavivirus structures differ in the amino acids that surround a glycosylation site in the virus shell. They wrote, “The structure of Zika virus is similar to other known flavivirus structures except for the ~10 amino acids that surround the Asn154 glycosylation site found in each of the 180 envelope glycoproteins that make up the icosahedral shell.”
The glycosylation site where Zika virus differs from other flaviviruses protrudes from the surface of the virus. A carbohydrate molecule consisting of various sugars is attached to the viral protein surface at this site. In other viruses it has been shown that as the virus projects a glycosylation site outward, an attachment receptor molecule on the surface of a human cell recognises the sugars and binds to them, noted Kuhn.
The glycosylation site and surrounding residues on Zika virus may also be involved in attachment to human cells, and the differences in the amino acids between different flaviviruses could signify differences in the kinds of molecules to which the virus can attach and the different human cells it can infect, Rossmann said.
“If this site functions as it does in dengue and is involved in attachment to human cells, it could be a good spot to target an antiviral compound,” Rossmann said. “If this is the case, perhaps an inhibitor could be designed to block this function and keep the virus from attaching to and infecting human cells.”
The team plans to pursue further testing to evaluate the different regions as targets for treatment and to develop potential therapeutic molecules, said Kuhn.
Giant squid found hidden in submarine canyons off Western Australia
Using eDNA, Australian scientists have found hidden biodiversity off the WA coast, including the...
Could this 'PFAS trap' remove the most difficult-to-capture variants from water?
Flinders University scientists have showcased the use of a nano-sized molecular cage that acts as...
What journalists expect from the scientists they speak to
Peer review is often treated as the end of the story, but for journalists it is usually the point...




