Lab-on-a-chip used to study single bacterial cells
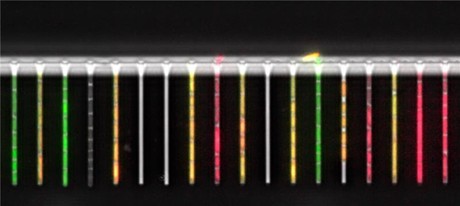
European researchers have set up a lab-on-a-chip, hardly bigger than a matchbox in size, which enables them to study gene regulation in single bacterial cells. Their work has been published in the journal Nature Communications.
Single bacterial cells grow in about 2000 channels of a thousandth of a millimetre in diameter and can be individually studied in detail by the researchers at the University of Basel, Switzerland. By recording thousands of microscopic images at short time intervals, the precise growth and behaviour of many generations of individual E. coli bacteria can be tracked over several days.
The huge amount of raw data generated is automatically analysed and precisely quantified by the chip’s accompanying image analysis software, known as MoMA. The software was developed in collaboration with scientists at the Max Planck Institute of Molecular Cell Biology and Genetics in Dresden, Germany.
Using the new system, the researchers can now study precisely how genes are regulated in single cells under changing environmental conditions, enabling them to gain insights not only into gene regulatory processes but also the diversity of adaptive responses of bacteria to varying environments. For example, it is possible to investigate how individual bacterial cells respond to a sudden exposure to an antibiotic: whether they die, stop growing or simply continue to divide undisturbed. It is also possible to observe the antibiotic’s increasing effect duration on the cells, which helps us understand why antibiotics do not always kill all pathogens.
“With the microfluidic chip we can also answer how bacteria communicate with each other, how they respond to stress or whether the relationship of bacterial strains plays a role in adaptation strategies,” said Professor Erik van Nimwegen, head of the Basel research group. “Such single-cell analyses are very important, because measurements of entire cell communities are often misleading since all the heterogeneity of the single cells has been averaged out.”
The researchers demonstrated the efficiency of the chip laboratory using a model system of gene regulation, the lac operon. The lac operon of E. coli contains genes involved in lactose metabolism, and is expressed only when lactose is present and glucose is absent.
“We have used green fluorescent protein to observe how E. coli bacteria respond to alternating nutrient changes from glucose to lactose,” Professor van Nimwegen said. “The lac operon has been studied for more than 50 years, and still we discovered new important properties when looking at it with single-cell resolution.”
In the first round, the bacteria switched to lactose turnover with a time lag. However, repeated switching from glucose to lactose led to a much faster adaptation of the cells as they started growing much earlier.
“Surprisingly, the lag times are similar in genetically related cells, suggesting that bacteria retain a memory of the behaviour of their ancestors,” Professor van Nimwegen said.
Microplastics found to alter the human gut microbiome
Microplastic-treated cultures showed a consistent and significant increase in acidity (lower pH...
Sustainable, self-repairing, antimicrobial polymers developed
From medicine to electronics and optics, new materials developed by scientists at Kaunas...
A better way to create conductive polymers
New research disproves the longstanding belief that to create conductive polymers, substances...




